2.5. Step 4: Load Data¶
Warning
WIP
Step number 4 brings us to the core functionality of openEHR: creating and storing clinical data! For this purpose, we will re-use the Template that we created in step 1. As data instances in openEHR are stored as instances of its Reference Model, it’s rather difficult to read for humans. However, this level of abstraction is needed inside the backend to achieve high scalability.
For application developers, more accessible formats are needed. There are two options: The EHRBase Client Library and using Flat Forms.
2.5.1. EHRBase Client Library¶
The EHRbase Client Library allows to use a Template (as OPT) as input and automatically create java classes. These can then be used to create the data. We explain this processes step by step. We assume that you have successfully built the Client Library.
Firstly, you need to create the Java classes from the OPT. This could look like as follows:
java -jar client-library-0.2.0.jar -opt "C:\Users\MyUser\Desktop\HiGHmed_Cardio_Monitoring_v1.opt" -out "C:\openEHR SDK\ehrbase_client_library\src\test\java\org\ehrbase\client\classgenerator" -package ""
You should find a file named HiGHmed_Cardio_Monitoring_v1.java inside your project structure that should look like this:
...
@Entity
@Archetype("openEHR-EHR-COMPOSITION.self_monitoring.v0")
@Template("HiGHmed_Cardio_Monitoring.v1")
public class HighmedCardioMonitoringV1 {
@Path("/context/end_time|value")
private TemporalAccessor endTimeValue;
@Path("/language")
private CodePhrase language;
@Path("/context/health_care_facility")
private PartyIdentified healthCareFacility;
@Path("/composer|external_ref")
private PartyRef composerExternalref;
...
Next, we can create a new test function like this:
public static HighmedCardioMonitoringV1 buildCardioExample(){
//Create the composition instance and add metadata
HighmedCardioMonitoringV1 cardioMonitoring = new HighmedCardioMonitoringV1();
cardioMonitoring.setLanguage(new CodePhrase(new TerminologyId("ISO_639-1"), "de"));
cardioMonitoring.setTerritory(new CodePhrase(new TerminologyId("ISO_3166-1"), "DE"));
cardioMonitoring.setSettingDefiningcode(new CodePhrase(new TerminologyId("openehr"), "229"));
//Create a blood pressure object
HighmedCardioMonitoringV1.Blutdruck bloodpressure = new HighmedCardioMonitoringV1.Blutdruck();
//Add data for systolic and diastolic blood pressure
bloodpressure.setSystolischMagnitude(160d);
bloodpressure.setSystolischUnits("mm[HG]");
bloodpressure.setDiastolischMagnitude(120d);
bloodpressure.setDiastolischUnits("mm[HG]");
//Add data for a medical device
HighmedCardioMonitoringV1.Blutdruck.MedizingerT geraet = new HighmedCardioMonitoringV1.Blutdruck.MedizingerT();
geraet.setBeschreibungValue("OMRON Sensor");
DvIdentifier identifier = new DvIdentifier();
identifier.setId("4567879799");
geraet.setEindeutigeIdentifikationsnummerId(identifier);
List<HighmedCardioMonitoringV1.Blutdruck> bpList = new ArrayList<>();
bpList.add(bloodpressure);
cardioMonitoring.setBlutdruck(bpList);
return cardioMonitoring;
}
Finally, the composition can be sent to the openEHR server:
CompositionEndpoint compositionEndpoint = openEhrClient.compositionEndpoint(ehr);
UUID compositionId = compositionEndpoint.saveCompositionEntity(highmedCardioMonitoringV1);
2.5.2. Flat Format¶
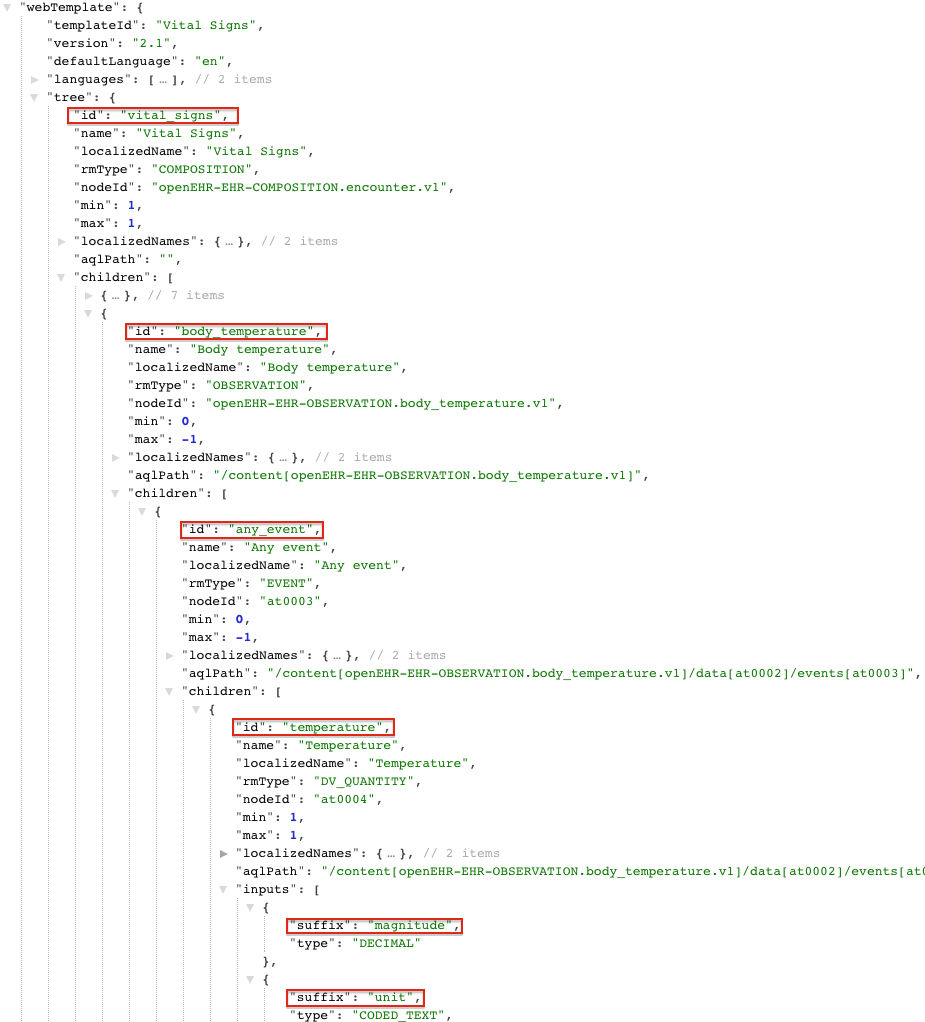
Another alternative to using the Client Library is to use a Simplified Data Template also known as the “Flat format”. In particular, we’ll be looking at the simplified IM Simplified Data template (simSDT) which is based on the web template format created by Marand for the Better platform. The first thing you need is to get the Web Template version of the Template. The ADL Designer tool allows you to export templates as Web Templates. An example of a simple Body Temperature Web Template (borrowed from EhrScape Examples) would look like this:
Next, we create the composition from the Web Template as a simple key-value pair with the keys being a path
obtained by concatenating the id of each level delimited by a /. The last segment is the suffix and uses | as a delimiter.
For example, in the above image all the id to be concatenated are highlighted in red.
So the paths built from the above example would look like:
vital_signs/body_temperature/any_event/temperature|magnitude
vital_signs/body_temperature/any_event/temperature|unit
The value of these above keys would be the actual data. Representing this in JSON would look like:
{
"vital_signs/body_temperature/any_event/temperature|magnitude": 92,
"vital_signs/body_temperature/any_event/temperature|unit": "°C"
}
However, since the cardinality of the body_temperature and any_event elements are -1 it means that
the composition can have an infinite number of body_temperature and body_temperature recorded in the same composition.
To resolve this, we have to index the path like so:
vital_signs/body_temperature:0/any_event:0/temperature|magnitude
vital_signs/body_temperature:0/any_event:0/temperature|unit
With these paths, and more context data, a composition with multiple recordings of body temperature will look like:
{
"ctx/time": "2014-03-19T13:10:00.000Z",
"ctx/language": "en",
"ctx/territory": "CA",
"vital_signs/body_temperature:0/any_event:0/time": "2014-03-19T13:10:00.000Z",
"vital_signs/body_temperature:0/any_event:0/temperature|magnitude": 37.1,
"vital_signs/body_temperature:0/any_event:0/temperature|unit": "°C",
"vital_signs/body_temperature:0/any_event:1/time": "2014-03-19T16:33:00.000Z",
"vital_signs/body_temperature:0/any_event:1/temperature|magnitude": 37.7,
"vital_signs/body_temperature:0/any_event:1/temperature|unit": "°C"
}
The API endpoints for the Flat Format is different from the normal composition API. More details can be found in this Postman Collection. To use the Flat Format, the latest version of EHRBase should be used. More information can be found here.
Congratulations, you stored your first clinical data inside EHRbase! Next, we will take a look how we can retrieve the data using the Archetype Query Language.